aLeaves - [èɪlíːvz]
first step to build zoologically informative phylogenetic treesBasic concept - why aLeaves?
The history of every gene in the human genome, as well as every gene that was lost before the establishment of the human genome, is traceable by collecting homologs of the gene of particular interest. This web server allows researchers and even non-researchers to explore protein-coding landscape of vertebrate genomes.
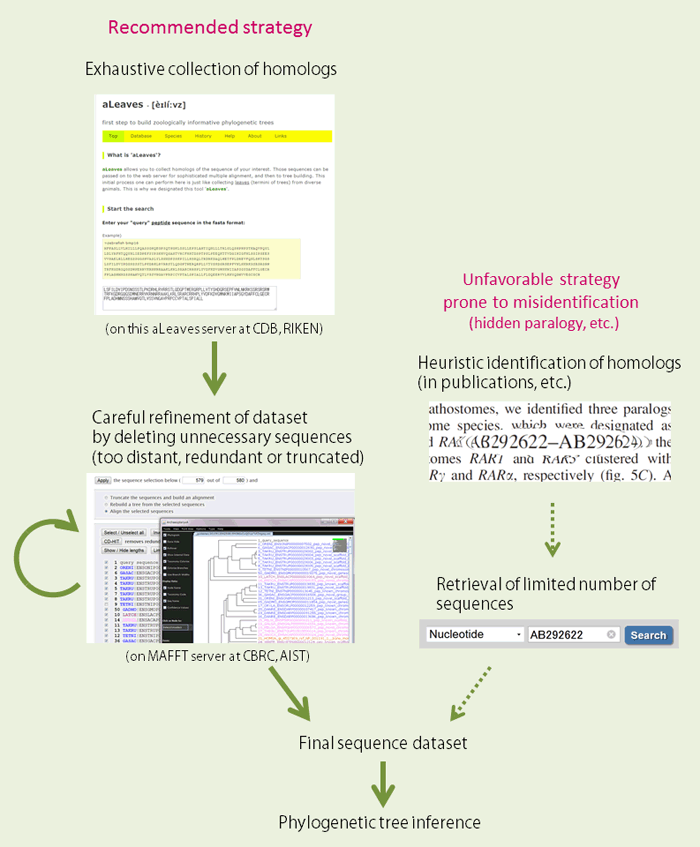
With all sequence information produced thanks to recent rapid innovation of sequencing technologies, typical work flow in which we handle the sequence information tends to be out-dated. By starting with exhaustive (but still not complete, of course) search of homologs, aLeaves provides a solution to avoid any misidentification in interpretation of molecular phylogeny (e.g., hidden paralogy).
About the developers
Acknowledgement
The aLeaves project is funded by RIKEN.
To cite aLeaves
The article introducing the aLeaves server has been published in the Web Server Issue 2013 of the journal Nucleic Acids Research.
aLeaves
facilitates on-demand exploration of metazoan gene family trees on
MAFFT sequence alignment server with enhanced interactivity.
Shigehiro Kuraku,
Christian Zmasek, Osamu Nishimura, and Kazutaka Katoh.
Nucleic Acids Research,
2013. 41 (W1): W22-W28.
Contact
For further info, please contact the chief administrator,
Shigehiro Kuraku (shigehiro.kuraku
at riken.jp).